Research
In general, our research interests center on phylogenetic comparative biology, functional morphology and biomechanics, and herpetology. We use data from museum specimens, lab experiments, and fieldwork to address questions of diversification, phenotypic evolution, and comparative biomechanics. Most of our work is collaborative, as we work with engineers, biomechanists, and systematists to greatly improve the range of questions we can ask (and hopefully answer!).
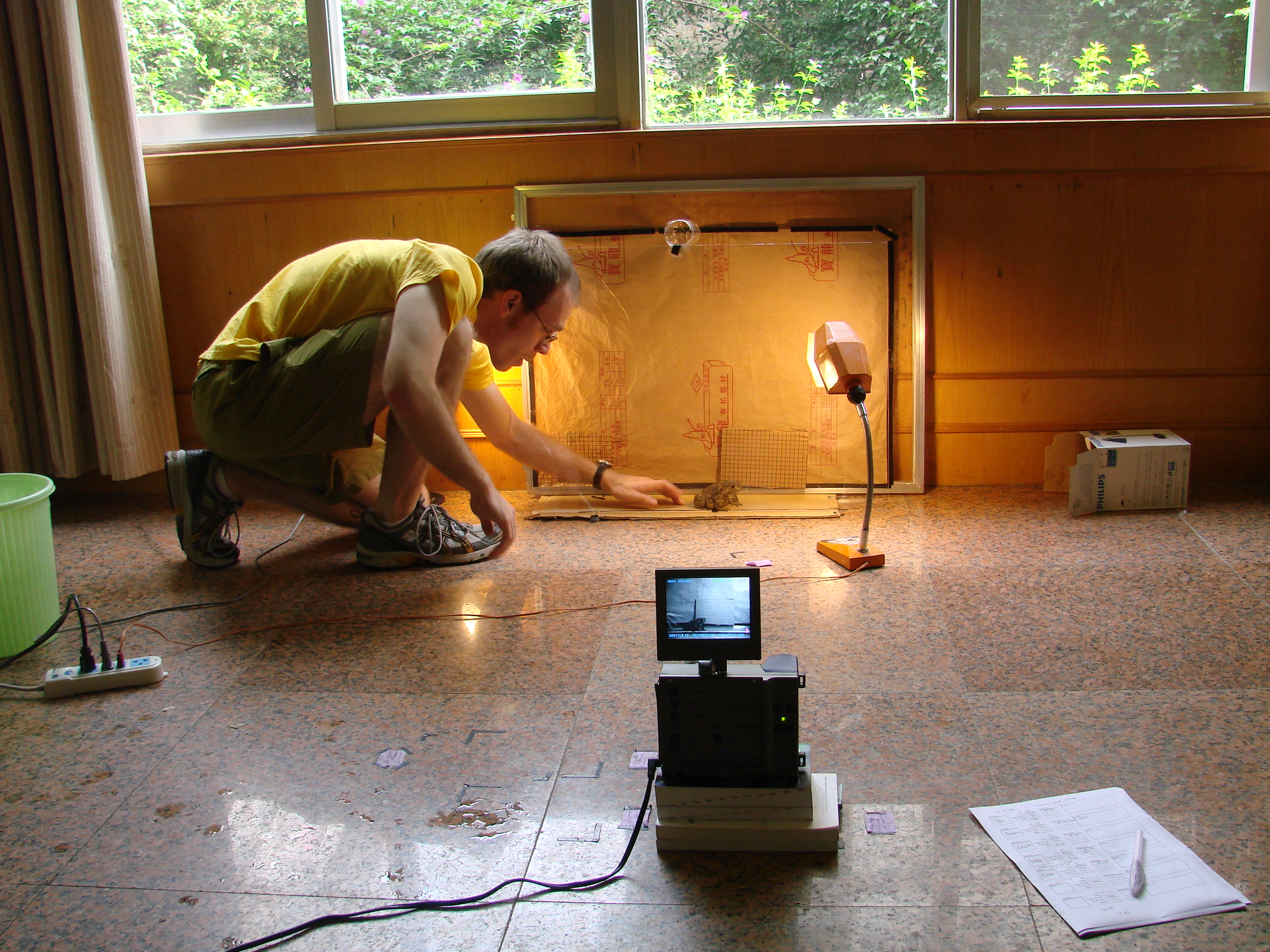 Evolutionary biomechanics
Evolutionary biomechanics
We are interested in the power of comparative approaches (i.e. studies across diverse species) in understanding the evolution of biomechanical systems. The questions we address in this part of our research are partly descriptive (“How does this work mechanistically? What is the diversity of mechanics in this behavior?”) and partly evolutionary (“How did this fantastic performance evolve? What are the consequences for macroevolution?”). We are currently trying to understand the mechanics and evolution of jumping, swimming, adhesion, and burrowing in frogs, using the power of phylogenetic comparative methods to examine variation one does not see within a single species and with the underlying question of “How has a fundamentally jumping organism evolved to do so many other things?” To answer this question and understand its consequences for the evolution of body form, we examine trade-offs, many-to-one mapping of form onto function, and modularity/integration.
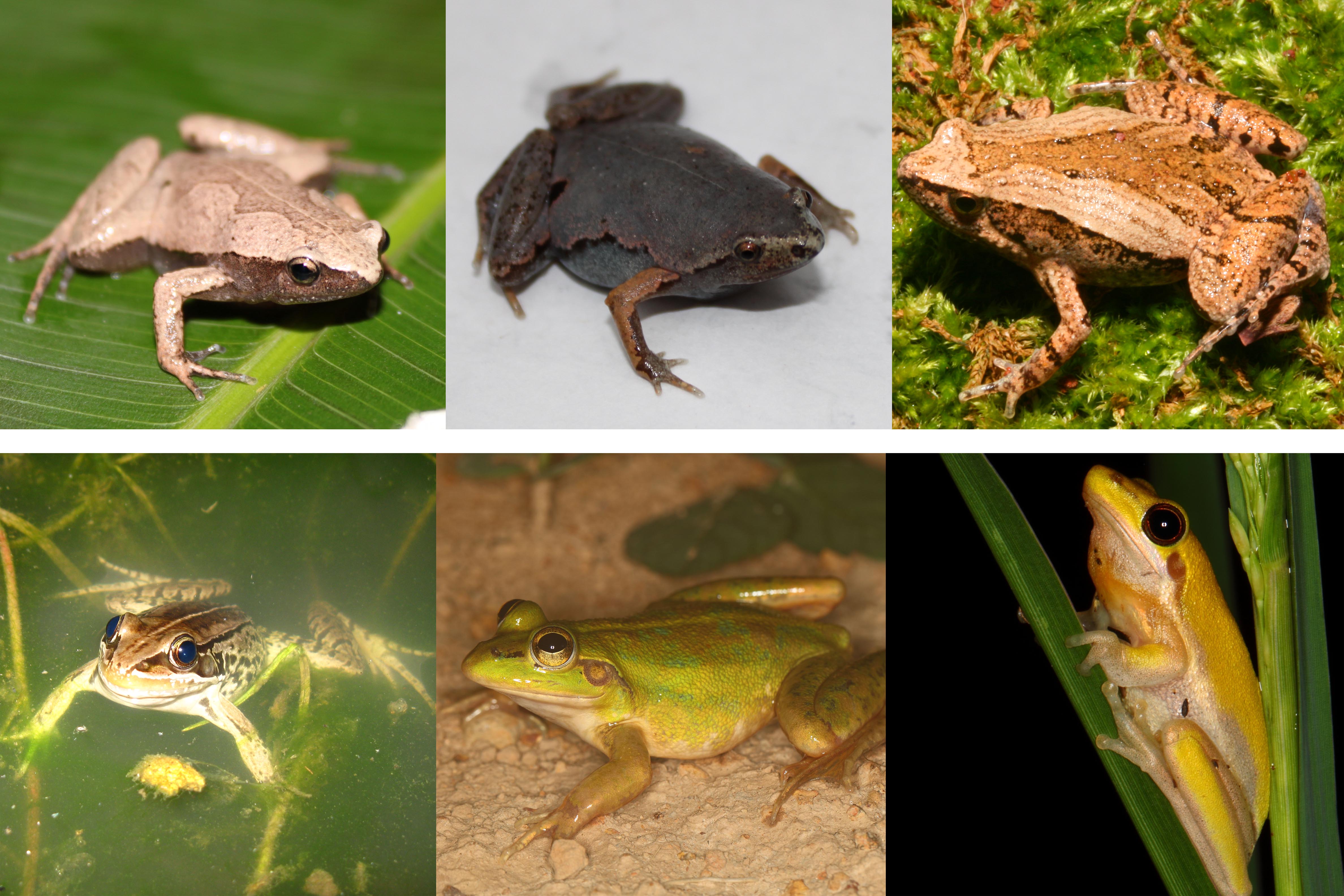 Ecomorphology, evolutionary history, and convergence
Ecomorphology, evolutionary history, and convergence
We are interested in asking whether (and how much) evolution is replicable – in the famous (paraphrased) words of Stephan J. Gould (1989, book Wonderful Life), if we were to replay the tape of evolution, would we always get the same answer (or different, as Gould surmised)? Given similar ecological circumstances, will roughly similar organisms (in our case, frogs) converge to the same adaptive solution? To address these questions, we collect morphological and functional data on museum specimens of frogs from around the world and from our own fieldwork. We then analyze these data to relate body form and function to ecology (primarily microhabitat), to test convergence, and to understand the various factors (from biogeography to biomechanics) affect the macroevolution of form, function, and ecology in frogs. The answers will help to understand why we see the spectacular (and odd, crazy, weird) ecologies and morphologies that so interested us (or at least me!) as kids and led us to science in the first place.
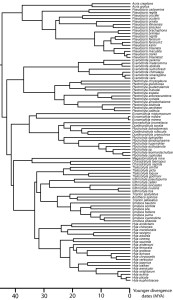 Causes of diverse patterns of phylogenetic diversification
Causes of diverse patterns of phylogenetic diversification
The phylogeny of any clade of extant organisms contains information about the diversification of the group – for example, rates of speciation, rates of extinction, and temporal variation in these rates. Some groups show constant rates of diversification, while others show most diversification either early in the clade’s history or much closer to the present. We are working to understand the causes of variation in amphibian and reptile diversification rates.
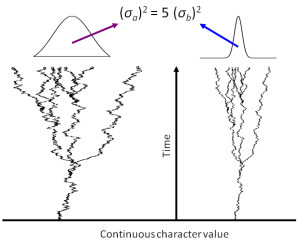 Phylogenetic, comparative, and statistical methods
Phylogenetic, comparative, and statistical methods
Phylogenetic, comparative, and general statistical methods also strongly interest us and we have developed novel analytical or simulation-based statistical tests for many of our papers. These include tests of community assembly, tests of evolutionary convergence, and resampling approaches to account for sampling artifacts. For this work, we extensively employ R and occasionally other tools (the bash command shell, MatLab, Python) to execute the tests we have developed, to run simulations, and to automate data extraction, formatting, and statistical analyses on large datasets (both our own and those of colleagues). Details of the methods can be found in Publications, particularly in the supplementary methods. You can find the code from our published papers here.